scipy.stats.multivariate_normal#
- scipy.stats.multivariate_normal = <scipy.stats._multivariate.multivariate_normal_gen object>[source]#
A multivariate normal random variable.
The mean keyword specifies the mean. The cov keyword specifies the covariance matrix.
- Parameters:
- meanarray_like, default:
[0] Mean of the distribution.
- covarray_like or
Covariance, default:[1] Symmetric positive (semi)definite covariance matrix of the distribution.
- allow_singularbool, default:
False Whether to allow a singular covariance matrix. This is ignored if cov is a
Covarianceobject.- seed{None, int, np.random.RandomState, np.random.Generator}, optional
Used for drawing random variates. If seed is None, the RandomState singleton is used. If seed is an int, a new
RandomStateinstance is used, seeded with seed. If seed is already aRandomStateorGeneratorinstance, then that object is used. Default is None.
- meanarray_like, default:
Methods
pdf(x, mean=None, cov=1, allow_singular=False)
Probability density function.
logpdf(x, mean=None, cov=1, allow_singular=False)
Log of the probability density function.
cdf(x, mean=None, cov=1, allow_singular=False, maxpts=1000000*dim, abseps=1e-5, releps=1e-5, lower_limit=None)
Cumulative distribution function.
logcdf(x, mean=None, cov=1, allow_singular=False, maxpts=1000000*dim, abseps=1e-5, releps=1e-5)
Log of the cumulative distribution function.
rvs(mean=None, cov=1, size=1, random_state=None)
Draw random samples from a multivariate normal distribution.
entropy(mean=None, cov=1)
Compute the differential entropy of the multivariate normal.
marginal(dimensions, mean=None, cov=1, allow_singular=False)
Return a marginal multivariate normal distribution.
fit(x, fix_mean=None, fix_cov=None)
Fit a multivariate normal distribution to data.
Notes
Setting the parameter mean to None is equivalent to having mean be the zero-vector. The parameter cov can be a scalar, in which case the covariance matrix is the identity times that value, a vector of diagonal entries for the covariance matrix, a two-dimensional array_like, or a
Covarianceobject.The covariance matrix cov may be an instance of a subclass of
Covariance, e.g. scipy.stats.CovViaPrecision. If so, allow_singular is ignored.Otherwise, cov must be a symmetric positive semidefinite matrix when allow_singular is True; it must be (strictly) positive definite when allow_singular is False. Symmetry is not checked; only the lower triangular portion is used. The determinant and inverse of cov are computed as the pseudo-determinant and pseudo-inverse, respectively, so that cov does not need to have full rank.
The probability density function for
multivariate_normalis\[f(x) = \frac{1}{\sqrt{(2 \pi)^k \det \Sigma}} \exp\left( -\frac{1}{2} (x - \mu)^T \Sigma^{-1} (x - \mu) \right),\]where \(\mu\) is the mean, \(\Sigma\) the covariance matrix, \(k\) the rank of \(\Sigma\). In case of singular \(\Sigma\), SciPy extends this definition according to [1].
Added in version 0.14.0.
References
[1]Multivariate Normal Distribution - Degenerate Case, Wikipedia, https://en.wikipedia.org/wiki/Multivariate_normal_distribution#Degenerate_case
Examples
>>> import numpy as np >>> import matplotlib.pyplot as plt >>> from scipy.stats import multivariate_normal
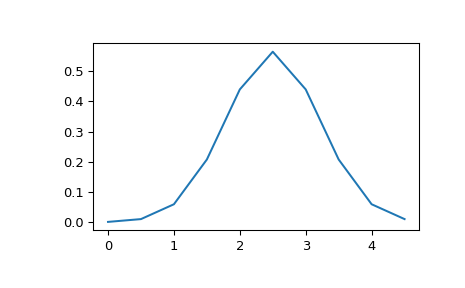
>>> x = np.linspace(0, 5, 10, endpoint=False) >>> y = multivariate_normal.pdf(x, mean=2.5, cov=0.5); y array([ 0.00108914, 0.01033349, 0.05946514, 0.20755375, 0.43939129, 0.56418958, 0.43939129, 0.20755375, 0.05946514, 0.01033349]) >>> fig1 = plt.figure() >>> ax = fig1.add_subplot(111) >>> ax.plot(x, y) >>> plt.show()
Alternatively, the object may be called (as a function) to fix the mean and covariance parameters, returning a “frozen” multivariate normal random variable:
>>> rv = multivariate_normal(mean=None, cov=1, allow_singular=False) >>> # Frozen object with the same methods but holding the given >>> # mean and covariance fixed.
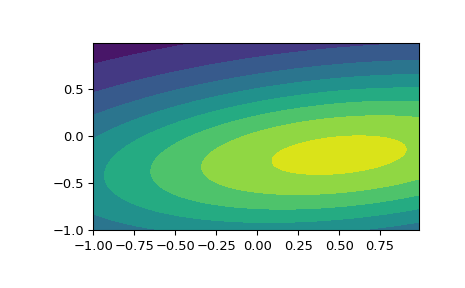
The input quantiles can be any shape of array, as long as the last axis labels the components. This allows us for instance to display the frozen pdf for a non-isotropic random variable in 2D as follows:
>>> x, y = np.mgrid[-1:1:.01, -1:1:.01] >>> pos = np.dstack((x, y)) >>> rv = multivariate_normal([0.5, -0.2], [[2.0, 0.3], [0.3, 0.5]]) >>> fig2 = plt.figure() >>> ax2 = fig2.add_subplot(111) >>> ax2.contourf(x, y, rv.pdf(pos))