differential_entropy#
- scipy.stats.differential_entropy(values, *, window_length=None, base=None, axis=0, method='auto', nan_policy='propagate', keepdims=False)[source]#
Given a sample of a distribution, estimate the differential entropy.
Several estimation methods are available using the method parameter. By default, a method is selected based the size of the sample.
- Parameters:
- valuessequence
Sample from a continuous distribution.
- window_lengthint, optional
Window length for computing Vasicek estimate. Must be an integer between 1 and half of the sample size. If
None(the default), it uses the heuristic value\[\left \lfloor \sqrt{n} + 0.5 \right \rfloor\]where \(n\) is the sample size. This heuristic was originally proposed in [2] and has become common in the literature.
- basefloat, optional
The logarithmic base to use, defaults to
e(natural logarithm).- axisint or None, default: 0
If an int, the axis of the input along which to compute the statistic. The statistic of each axis-slice (e.g. row) of the input will appear in a corresponding element of the output. If
None, the input will be raveled before computing the statistic.- method{‘vasicek’, ‘van es’, ‘ebrahimi’, ‘correa’, ‘auto’}, optional
The method used to estimate the differential entropy from the sample. Default is
'auto'. See Notes for more information.- nan_policy{‘propagate’, ‘omit’, ‘raise’}
Defines how to handle input NaNs.
propagate: if a NaN is present in the axis slice (e.g. row) along which the statistic is computed, the corresponding entry of the output will be NaN.omit: NaNs will be omitted when performing the calculation. If insufficient data remains in the axis slice along which the statistic is computed, the corresponding entry of the output will be NaN.raise: if a NaN is present, aValueErrorwill be raised.
- keepdimsbool, default: False
If this is set to True, the axes which are reduced are left in the result as dimensions with size one. With this option, the result will broadcast correctly against the input array.
- Returns:
- entropyfloat
The calculated differential entropy.
Notes
This function will converge to the true differential entropy in the limit
\[n \to \infty, \quad m \to \infty, \quad \frac{m}{n} \to 0\]The optimal choice of
window_lengthfor a given sample size depends on the (unknown) distribution. Typically, the smoother the density of the distribution, the larger the optimal value ofwindow_length[1].The following options are available for the method parameter.
'vasicek'uses the estimator presented in [1]. This is one of the first and most influential estimators of differential entropy.'van es'uses the bias-corrected estimator presented in [3], which is not only consistent but, under some conditions, asymptotically normal.'ebrahimi'uses an estimator presented in [4], which was shown in simulation to have smaller bias and mean squared error than the Vasicek estimator.'correa'uses the estimator presented in [5] based on local linear regression. In a simulation study, it had consistently smaller mean square error than the Vasiceck estimator, but it is more expensive to compute.'auto'selects the method automatically (default). Currently, this selects'van es'for very small samples (<10),'ebrahimi'for moderate sample sizes (11-1000), and'vasicek'for larger samples, but this behavior is subject to change in future versions.
All estimators are implemented as described in [6].
Beginning in SciPy 1.9,
np.matrixinputs (not recommended for new code) are converted tonp.ndarraybefore the calculation is performed. In this case, the output will be a scalar ornp.ndarrayof appropriate shape rather than a 2Dnp.matrix. Similarly, while masked elements of masked arrays are ignored, the output will be a scalar ornp.ndarrayrather than a masked array withmask=False.Array API Standard Support
differential_entropyhas experimental support for Python Array API Standard compatible backends in addition to NumPy. Please consider testing these features by setting an environment variableSCIPY_ARRAY_API=1and providing CuPy, PyTorch, JAX, or Dask arrays as array arguments. The following combinations of backend and device (or other capability) are supported.Library
CPU
GPU
NumPy
✅
n/a
CuPy
n/a
✅
PyTorch
✅
✅
JAX
✅
✅
Dask
✅
n/a
See Support for the array API standard for more information.
References
[1] (1,2)Vasicek, O. (1976). A test for normality based on sample entropy. Journal of the Royal Statistical Society: Series B (Methodological), 38(1), 54-59.
[2]Crzcgorzewski, P., & Wirczorkowski, R. (1999). Entropy-based goodness-of-fit test for exponentiality. Communications in Statistics-Theory and Methods, 28(5), 1183-1202.
[3]Van Es, B. (1992). Estimating functionals related to a density by a class of statistics based on spacings. Scandinavian Journal of Statistics, 61-72.
[4]Ebrahimi, N., Pflughoeft, K., & Soofi, E. S. (1994). Two measures of sample entropy. Statistics & Probability Letters, 20(3), 225-234.
[5]Correa, J. C. (1995). A new estimator of entropy. Communications in Statistics-Theory and Methods, 24(10), 2439-2449.
[6]Noughabi, H. A. (2015). Entropy Estimation Using Numerical Methods. Annals of Data Science, 2(2), 231-241. https://link.springer.com/article/10.1007/s40745-015-0045-9
Examples
>>> import numpy as np >>> from scipy.stats import differential_entropy, norm
Entropy of a standard normal distribution:
>>> rng = np.random.default_rng() >>> values = rng.standard_normal(100) >>> differential_entropy(values) 1.3407817436640392
Compare with the true entropy:
>>> float(norm.entropy()) 1.4189385332046727
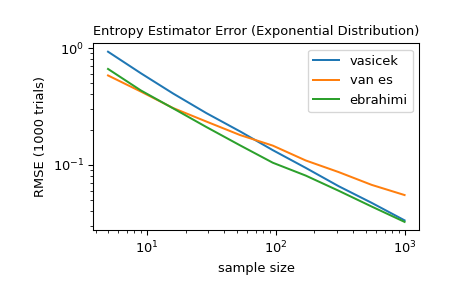
For several sample sizes between 5 and 1000, compare the accuracy of the
'vasicek','van es', and'ebrahimi'methods. Specifically, compare the root mean squared error (over 1000 trials) between the estimate and the true differential entropy of the distribution.>>> from scipy import stats >>> import matplotlib.pyplot as plt >>> >>> >>> def rmse(res, expected): ... '''Root mean squared error''' ... return np.sqrt(np.mean((res - expected)**2)) >>> >>> >>> a, b = np.log10(5), np.log10(1000) >>> ns = np.round(np.logspace(a, b, 10)).astype(int) >>> reps = 1000 # number of repetitions for each sample size >>> expected = stats.expon.entropy() >>> >>> method_errors = {'vasicek': [], 'van es': [], 'ebrahimi': []} >>> for method in method_errors: ... for n in ns: ... rvs = stats.expon.rvs(size=(reps, n), random_state=rng) ... res = stats.differential_entropy(rvs, method=method, axis=-1) ... error = rmse(res, expected) ... method_errors[method].append(error) >>> >>> for method, errors in method_errors.items(): ... plt.loglog(ns, errors, label=method) >>> >>> plt.legend() >>> plt.xlabel('sample size') >>> plt.ylabel('RMSE (1000 trials)') >>> plt.title('Entropy Estimator Error (Exponential Distribution)')