kmeans2#
- scipy.cluster.vq.kmeans2(data, k, iter=10, thresh=1e-05, minit='random', missing='warn', check_finite=True, *, rng=None, seed=None)[source]#
Classify a set of observations into k clusters using the k-means algorithm.
The algorithm attempts to minimize the Euclidean distance between observations and centroids. Several initialization methods are included.
- Parameters:
- datandarray
A ‘M’ by ‘N’ array of ‘M’ observations in ‘N’ dimensions or a length ‘M’ array of ‘M’ 1-D observations.
- kint or ndarray
The number of clusters to form as well as the number of centroids to generate. If minit initialization string is ‘matrix’, or if a ndarray is given instead, it is interpreted as initial cluster to use instead.
- iterint, optional
Number of iterations of the k-means algorithm to run. Note that this differs in meaning from the iters parameter to the kmeans function.
- threshfloat, optional
(not used yet)
- minitstr, optional
Method for initialization. Available methods are ‘random’, ‘points’, ‘++’ and ‘matrix’:
‘random’: generate k centroids from a Gaussian with mean and variance estimated from the data.
‘points’: choose k observations (rows) at random from data for the initial centroids.
‘++’: choose k observations accordingly to the kmeans++ method (careful seeding)
‘matrix’: interpret the k parameter as a k by M (or length k array for 1-D data) array of initial centroids.
- missingstr, optional
Method to deal with empty clusters. Available methods are ‘warn’ and ‘raise’:
‘warn’: give a warning and continue.
‘raise’: raise an ClusterError and terminate the algorithm.
- check_finitebool, optional
Whether to check that the input matrices contain only finite numbers. Disabling may give a performance gain, but may result in problems (crashes, non-termination) if the inputs do contain infinities or NaNs. Default: True
- rng{None, int,
numpy.random.Generator}, optional If rng is passed by keyword, types other than
numpy.random.Generatorare passed tonumpy.random.default_rngto instantiate aGenerator. If rng is already aGeneratorinstance, then the provided instance is used. Specify rng for repeatable function behavior.If this argument is passed by position or seed is passed by keyword, legacy behavior for the argument seed applies:
If seed is None (or
numpy.random), thenumpy.random.RandomStatesingleton is used.If seed is an int, a new
RandomStateinstance is used, seeded with seed.If seed is already a
GeneratororRandomStateinstance then that instance is used.
Changed in version 1.15.0: As part of the SPEC-007 transition from use of
numpy.random.RandomStatetonumpy.random.Generator, this keyword was changed from seed to rng. For an interim period, both keywords will continue to work, although only one may be specified at a time. After the interim period, function calls using the seed keyword will emit warnings. The behavior of both seed and rng are outlined above, but only the rng keyword should be used in new code.
- Returns:
- centroidndarray
A ‘k’ by ‘N’ array of centroids found at the last iteration of k-means.
- labelndarray
label[i] is the code or index of the centroid the ith observation is closest to.
See also
Notes
Array API Standard Support
kmeans2has experimental support for Python Array API Standard compatible backends in addition to NumPy. Please consider testing these features by setting an environment variableSCIPY_ARRAY_API=1and providing CuPy, PyTorch, JAX, or Dask arrays as array arguments. The following combinations of backend and device (or other capability) are supported.Library
CPU
GPU
NumPy
✅
n/a
CuPy
n/a
⛔
PyTorch
✅
⛔
JAX
⚠️ no JIT
⛔
Dask
⚠️ computes graph
n/a
See Support for the array API standard for more information.
References
[1]D. Arthur and S. Vassilvitskii, “k-means++: the advantages of careful seeding”, Proceedings of the Eighteenth Annual ACM-SIAM Symposium on Discrete Algorithms, 2007.
Examples
>>> from scipy.cluster.vq import kmeans2 >>> import matplotlib.pyplot as plt >>> import numpy as np
Create z, an array with shape (100, 2) containing a mixture of samples from three multivariate normal distributions.
>>> rng = np.random.default_rng() >>> a = rng.multivariate_normal([0, 6], [[2, 1], [1, 1.5]], size=45) >>> b = rng.multivariate_normal([2, 0], [[1, -1], [-1, 3]], size=30) >>> c = rng.multivariate_normal([6, 4], [[5, 0], [0, 1.2]], size=25) >>> z = np.concatenate((a, b, c)) >>> rng.shuffle(z)
Compute three clusters.
>>> centroid, label = kmeans2(z, 3, minit='points') >>> centroid array([[ 2.22274463, -0.61666946], # may vary [ 0.54069047, 5.86541444], [ 6.73846769, 4.01991898]])
How many points are in each cluster?
>>> counts = np.bincount(label) >>> counts array([29, 51, 20]) # may vary
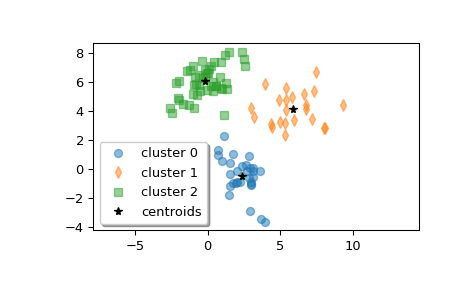
Plot the clusters.
>>> w0 = z[label == 0] >>> w1 = z[label == 1] >>> w2 = z[label == 2] >>> plt.plot(w0[:, 0], w0[:, 1], 'o', alpha=0.5, label='cluster 0') >>> plt.plot(w1[:, 0], w1[:, 1], 'd', alpha=0.5, label='cluster 1') >>> plt.plot(w2[:, 0], w2[:, 1], 's', alpha=0.5, label='cluster 2') >>> plt.plot(centroid[:, 0], centroid[:, 1], 'k*', label='centroids') >>> plt.axis('equal') >>> plt.legend(shadow=True) >>> plt.show()