Probability distributions#
SciPy has two infrastructures for working with probability distributions. This tutorial is for the older one, which has many pre-defined distributions; however, the new infrastructure can be used with most of these and has many advantages. For the new infrastructure, see Random Variable Transition Guide.
There are two general distribution classes that have been implemented for encapsulating continuous random variables and discrete random variables. Over 100 continuous random variables (RVs) and 20 discrete random variables have been implemented using these classes. For mathematical reference information about individual distributions, please see Continuous Statistical Distributions and Discrete Statistical Distributions.
All of the statistics functions are located in the sub-package
scipy.stats and a fairly complete listing of these functions and random
variables available can also be obtained from the docstring for the
stats sub-package.
In the discussion below, we mostly focus on continuous RVs. Nearly everything also applies to discrete variables, but we point out some differences here: Specific points for discrete distributions.
In the code samples below, we assume that the scipy.stats package
is imported as
>>> from scipy import stats
and in some cases we assume that individual objects are imported as
>>> from scipy.stats import norm
All distributions
Getting help#
First of all, all distributions are accompanied with help
functions. To obtain just some basic information, we print the relevant
docstring: print(stats.norm.__doc__).
To find the support, i.e., upper and lower bounds of the distribution, call:
>>> print('bounds of distribution lower: %s, upper: %s' % norm.support())
bounds of distribution lower: -inf, upper: inf
We can list all methods and properties of the distribution with
dir(norm). As it turns out, some of the methods are private,
although they are not named as such (their names do not start
with a leading underscore), for example veccdf, are only available
for internal calculation (those methods will give warnings when one tries to
use them, and will be removed at some point).
To obtain the real main methods, we list the methods of the frozen distribution. (We explain the meaning of a frozen distribution below).
>>> rv = norm()
>>> dir(rv) # reformatted
['__class__', '__delattr__', '__dict__', '__dir__', '__doc__', '__eq__',
'__format__', '__ge__', '__getattribute__', '__gt__', '__hash__',
'__init__', '__le__', '__lt__', '__module__', '__ne__', '__new__',
'__reduce__', '__reduce_ex__', '__repr__', '__setattr__', '__sizeof__',
'__str__', '__subclasshook__', '__weakref__', 'a', 'args', 'b', 'cdf',
'dist', 'entropy', 'expect', 'interval', 'isf', 'kwds', 'logcdf',
'logpdf', 'logpmf', 'logsf', 'mean', 'median', 'moment', 'pdf', 'pmf',
'ppf', 'random_state', 'rvs', 'sf', 'stats', 'std', 'var']
Finally, we can obtain the list of available distribution through introspection:
>>> dist_continu = [d for d in dir(stats) if
... isinstance(getattr(stats, d), stats.rv_continuous)]
>>> dist_discrete = [d for d in dir(stats) if
... isinstance(getattr(stats, d), stats.rv_discrete)]
>>> print('number of continuous distributions: %d' % len(dist_continu))
number of continuous distributions: 109
>>> print('number of discrete distributions: %d' % len(dist_discrete))
number of discrete distributions: 21
Common methods#
The main public methods for continuous RVs are:
rvs: Random Variates
pdf: Probability Density Function
cdf: Cumulative Distribution Function
sf: Survival Function (1-CDF)
ppf: Percent Point Function (Inverse of CDF)
isf: Inverse Survival Function (Inverse of SF)
stats: Return mean, variance, (Fisher’s) skew, or (Fisher’s) kurtosis
moment: non-central moments of the distribution
Let’s take a normal RV as an example.
>>> norm.cdf(0)
0.5
To compute the cdf at a number of points, we can pass a list or a numpy array.
>>> norm.cdf([-1., 0, 1])
array([ 0.15865525, 0.5, 0.84134475])
>>> import numpy as np
>>> norm.cdf(np.array([-1., 0, 1]))
array([ 0.15865525, 0.5, 0.84134475])
Thus, the basic methods, such as pdf, cdf, and so on, are vectorized.
Other generally useful methods are supported too:
>>> norm.mean(), norm.std(), norm.var()
(0.0, 1.0, 1.0)
>>> norm.stats(moments="mv")
(array(0.0), array(1.0))
To find the median of a distribution, we can use the percent point
function ppf, which is the inverse of the cdf:
>>> norm.ppf(0.5)
0.0
To generate a sequence of random variates, use the size keyword
argument:
>>> norm.rvs(size=3)
array([-0.35687759, 1.34347647, -0.11710531]) # random
Don’t think that norm.rvs(5) generates 5 variates:
>>> norm.rvs(5)
5.471435163732493 # random
Here, 5 with no keyword is being interpreted as the first possible
keyword argument, loc, which is the first of a pair of keyword arguments
taken by all continuous distributions.
This brings us to the topic of the next subsection.
Random number generation#
Drawing random numbers relies on generators from numpy.random package.
In the examples above, the specific stream of
random numbers is not reproducible across runs. To achieve reproducibility,
you can explicitly seed a random number generator. In NumPy, a generator
is an instance of numpy.random.Generator. Here is the canonical way to create
a generator:
>>> from numpy.random import default_rng
>>> rng = default_rng()
And fixing the seed can be done like this:
>>> # do NOT copy this value
>>> rng = default_rng(301439351238479871608357552876690613766)
Warning
Do not use this number or common values such as 0. Using just a
small set of seeds to instantiate larger state spaces means that
there are some initial states that are impossible to reach. This
creates some biases if everyone uses such values. A good way to
get a seed is to use a numpy.random.SeedSequence:
>>> from numpy.random import SeedSequence
>>> print(SeedSequence().entropy)
301439351238479871608357552876690613766 # random
The random_state parameter in distributions accepts an instance of
numpy.random.Generator class, or an integer, which is then used to
seed an internal Generator object:
>>> norm.rvs(size=5, random_state=rng)
array([ 0.47143516, -1.19097569, 1.43270697, -0.3126519 , -0.72058873]) # random
For further info, see NumPy’s documentation.
To learn more about the random number samplers implemented in SciPy, see non-uniform random number sampling tutorial and quasi monte carlo tutorial
Shifting and scaling#
All continuous distributions take loc and scale as keyword
parameters to adjust the location and scale of the distribution,
e.g., for the standard normal distribution, the location is the mean and
the scale is the standard deviation.
>>> norm.stats(loc=3, scale=4, moments="mv")
(array(3.0), array(16.0))
In many cases, the standardized distribution for a random variable X
is obtained through the transformation (X - loc) / scale. The
default values are loc = 0 and scale = 1.
Smart use of loc and scale can help modify the standard
distributions in many ways. To illustrate the scaling further, the
cdf of an exponentially distributed RV with mean \(1/\lambda\)
is given by
By applying the scaling rule above, it can be seen that by
taking scale = 1./lambda we get the proper scale.
>>> from scipy.stats import expon
>>> expon.mean(scale=3.)
3.0
Note
Distributions that take shape parameters may
require more than simple application of loc and/or
scale to achieve the desired form. For example, the
distribution of 2-D vector lengths given a constant vector
of length \(R\) perturbed by independent N(0, \(\sigma^2\))
deviations in each component is
rice(\(R/\sigma\), scale= \(\sigma\)). The first argument
is a shape parameter that needs to be scaled along with \(x\).
The uniform distribution is also interesting:
>>> from scipy.stats import uniform
>>> uniform.cdf([0, 1, 2, 3, 4, 5], loc=1, scale=4)
array([ 0. , 0. , 0.25, 0.5 , 0.75, 1. ])
Finally, recall from the previous paragraph that we are left with the
problem of the meaning of norm.rvs(5). As it turns out, calling a
distribution like this, the first argument, i.e., the 5, gets passed
to set the loc parameter. Let’s see:
>>> np.mean(norm.rvs(5, size=500))
5.0098355106969992 # random
Thus, to explain the output of the example of the last section:
norm.rvs(5) generates a single normally distributed random variate with
mean loc=5, because of the default size=1.
We recommend that you set loc and scale parameters explicitly, by
passing the values as keywords rather than as arguments. Repetition
can be minimized when calling more than one method of a given RV by
using the technique of Freezing a Distribution, as explained below.
Shape parameters#
While a general continuous random variable can be shifted and scaled
with the loc and scale parameters, some distributions require
additional shape parameters. For instance, the gamma distribution with density
requires the shape parameter \(a\). Observe that setting
\(\lambda\) can be obtained by setting the scale keyword to
\(1/\lambda\).
Let’s check the number and name of the shape parameters of the gamma distribution. (We know from the above that this should be 1.)
>>> from scipy.stats import gamma
>>> gamma.numargs
1
>>> gamma.shapes
'a'
Now, we set the value of the shape variable to 1 to obtain the exponential distribution, so that we compare easily whether we get the results we expect.
>>> gamma(1, scale=2.).stats(moments="mv")
(array(2.0), array(4.0))
Notice that we can also specify shape parameters as keywords:
>>> gamma(a=1, scale=2.).stats(moments="mv")
(array(2.0), array(4.0))
Freezing a distribution#
Passing the loc and scale keywords time and again can become
quite bothersome. The concept of freezing a RV is used to
solve such problems.
>>> rv = gamma(1, scale=2.)
By using rv we no longer have to include the scale or the shape
parameters anymore. Thus, distributions can be used in one of two
ways, either by passing all distribution parameters to each method
call (such as we did earlier) or by freezing the parameters for the
instance of the distribution. Let us check this:
>>> rv.mean(), rv.std()
(2.0, 2.0)
This is, indeed, what we should get.
Broadcasting#
The basic methods pdf, and so on, satisfy the usual numpy broadcasting rules. For
example, we can calculate the critical values for the upper tail of
the t distribution for different probabilities and degrees of freedom.
>>> stats.t.isf([0.1, 0.05, 0.01], [[10], [11]])
array([[ 1.37218364, 1.81246112, 2.76376946],
[ 1.36343032, 1.79588482, 2.71807918]])
Here, the first row contains the critical values for 10 degrees of freedom
and the second row for 11 degrees of freedom (d.o.f.). Thus, the
broadcasting rules give the same result of calling isf twice:
>>> stats.t.isf([0.1, 0.05, 0.01], 10)
array([ 1.37218364, 1.81246112, 2.76376946])
>>> stats.t.isf([0.1, 0.05, 0.01], 11)
array([ 1.36343032, 1.79588482, 2.71807918])
If the array with probabilities, i.e., [0.1, 0.05, 0.01] and the
array of degrees of freedom i.e., [10, 11, 12], have the same
array shape, then element-wise matching is used. As an example, we can
obtain the 10% tail for 10 d.o.f., the 5% tail for 11 d.o.f. and the
1% tail for 12 d.o.f. by calling
>>> stats.t.isf([0.1, 0.05, 0.01], [10, 11, 12])
array([ 1.37218364, 1.79588482, 2.68099799])
Specific points for discrete distributions#
Discrete distributions have mostly the same basic methods as the
continuous distributions. However pdf is replaced by the probability
mass function pmf, no estimation methods, such as fit, are
available, and scale is not a valid keyword parameter. The
location parameter, keyword loc, can still be used to shift the
distribution.
The computation of the cdf requires some extra attention. In the case of continuous distribution, the cumulative distribution function is, in most standard cases, strictly monotonic increasing in the bounds (a,b) and has, therefore, a unique inverse. The cdf of a discrete distribution, however, is a step function, hence the inverse cdf, i.e., the percent point function, requires a different definition:
ppf(q) = min{x : cdf(x) >= q, x integer}
For further info, see the docs here.
We can look at the hypergeometric distribution as an example
>>> from scipy.stats import hypergeom
>>> [M, n, N] = [20, 7, 12]
If we use the cdf at some integer points and then evaluate the ppf at those cdf values, we get the initial integers back, for example
>>> x = np.arange(4) * 2
>>> x
array([0, 2, 4, 6])
>>> prb = hypergeom.cdf(x, M, n, N)
>>> prb
array([ 1.03199174e-04, 5.21155831e-02, 6.08359133e-01,
9.89783282e-01])
>>> hypergeom.ppf(prb, M, n, N)
array([ 0., 2., 4., 6.])
If we use values that are not at the kinks of the cdf step function, we get the next higher integer back:
>>> hypergeom.ppf(prb + 1e-8, M, n, N)
array([ 1., 3., 5., 7.])
>>> hypergeom.ppf(prb - 1e-8, M, n, N)
array([ 0., 2., 4., 6.])
Fitting distributions#
The main additional methods of the not frozen distribution are related to the estimation of distribution parameters:
- fit: maximum likelihood estimation of distribution parameters, including location
and scale
fit_loc_scale: estimation of location and scale when shape parameters are given
nnlf: negative log likelihood function
expect: calculate the expectation of a function against the pdf or pmf
Performance issues and cautionary remarks#
The performance of the individual methods, in terms of speed, varies widely by distribution and method. The results of a method are obtained in one of two ways: either by explicit calculation, or by a generic algorithm that is independent of the specific distribution.
Explicit calculation, on the one hand, requires that the method is
directly specified for the given distribution, either through analytic
formulas or through special functions in scipy.special or
numpy.random for rvs. These are usually relatively fast
calculations.
The generic methods, on the other hand, are used if the distribution
does not specify any explicit calculation. To define a distribution,
only one of pdf or cdf is necessary; all other methods can be derived
using numeric integration and root finding. However, these indirect
methods can be very slow. As an example, rgh =
stats.gausshyper.rvs(0.5, 2, 2, 2, size=100) creates random
variables in a very indirect way and takes about 19 seconds for 100
random variables on my computer, while one million random variables
from the standard normal or from the t distribution take just above
one second.
Remaining issues#
The distributions in scipy.stats have recently been corrected and improved
and gained a considerable test suite; however, a few issues remain:
The distributions have been tested over some range of parameters; however, in some corner ranges, a few incorrect results may remain.
The maximum likelihood estimation in fit does not work with default starting parameters for all distributions and the user needs to supply good starting parameters. Also, for some distribution using a maximum likelihood estimator might inherently not be the best choice.
Building specific distributions#
The next examples shows how to build your own distributions. Further examples show the usage of the distributions and some statistical tests.
Making a continuous distribution, i.e., subclassing rv_continuous#
Making continuous distributions is fairly simple.
>>> from scipy import stats
>>> class deterministic_gen(stats.rv_continuous):
... def _cdf(self, x):
... return np.where(x < 0, 0., 1.)
... def _stats(self):
... return 0., 0., 0., 0.
>>> deterministic = deterministic_gen(name="deterministic")
>>> deterministic.cdf(np.arange(-3, 3, 0.5))
array([ 0., 0., 0., 0., 0., 0., 1., 1., 1., 1., 1., 1.])
Interestingly, the pdf is now computed automatically:
>>> deterministic.pdf(np.arange(-3, 3, 0.5))
array([ 0.00000000e+00, 0.00000000e+00, 0.00000000e+00,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00,
5.83333333e+04, 4.16333634e-12, 4.16333634e-12,
4.16333634e-12, 4.16333634e-12, 4.16333634e-12])
Be aware of the performance issues mentioned in Performance issues and cautionary remarks. The computation of unspecified common methods can become very slow, since only general methods are called, which, by their very nature, cannot use any specific information about the distribution. Thus, as a cautionary example:
>>> from scipy.integrate import quad
>>> quad(deterministic.pdf, -1e-1, 1e-1)
(4.163336342344337e-13, 0.0)
But this is not correct: the integral over this pdf should be 1. Let’s make the integration interval smaller:
>>> quad(deterministic.pdf, -1e-3, 1e-3) # warning removed
(1.000076872229173, 0.0010625571718182458)
This looks better. However, the problem originated from the fact that the pdf is not specified in the class definition of the deterministic distribution.
Subclassing rv_discrete#
In the following, we use stats.rv_discrete to generate a discrete distribution that has the probabilities of the truncated normal for the intervals centered around the integers.
General info
From the docstring of rv_discrete, help(stats.rv_discrete),
“You can construct an arbitrary discrete rv where P{X=xk} = pk by passing to the rv_discrete initialization method (through the values= keyword) a tuple of sequences (xk, pk) which describes only those values of X (xk) that occur with nonzero probability (pk).”
Next to this, there are some further requirements for this approach to work:
The keyword name is required.
The support points of the distribution xk have to be integers.
The number of significant digits (decimals) needs to be specified.
In fact, if the last two requirements are not satisfied, an exception may be raised or the resulting numbers may be incorrect.
An example
Let’s do the work. First:
>>> npoints = 20 # number of integer support points of the distribution minus 1
>>> npointsh = npoints // 2
>>> npointsf = float(npoints)
>>> nbound = 4 # bounds for the truncated normal
>>> normbound = (1+1/npointsf) * nbound # actual bounds of truncated normal
>>> grid = np.arange(-npointsh, npointsh+2, 1) # integer grid
>>> gridlimitsnorm = (grid-0.5) / npointsh * nbound # bin limits for the truncnorm
>>> gridlimits = grid - 0.5 # used later in the analysis
>>> grid = grid[:-1]
>>> probs = np.diff(stats.truncnorm.cdf(gridlimitsnorm, -normbound, normbound))
>>> gridint = grid
And, finally, we can subclass rv_discrete:
>>> normdiscrete = stats.rv_discrete(values=(gridint,
... np.round(probs, decimals=7)), name='normdiscrete')
Now that we have defined the distribution, we have access to all common methods of discrete distributions.
>>> print('mean = %6.4f, variance = %6.4f, skew = %6.4f, kurtosis = %6.4f' %
... normdiscrete.stats(moments='mvsk'))
mean = -0.0000, variance = 6.3302, skew = 0.0000, kurtosis = -0.0076
>>> nd_std = np.sqrt(normdiscrete.stats(moments='v'))
Testing the implementation
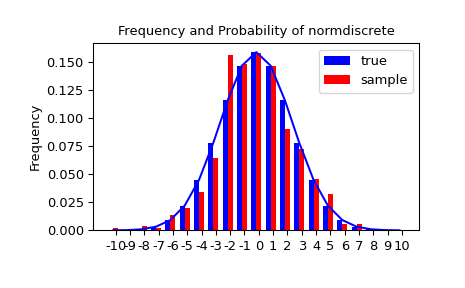
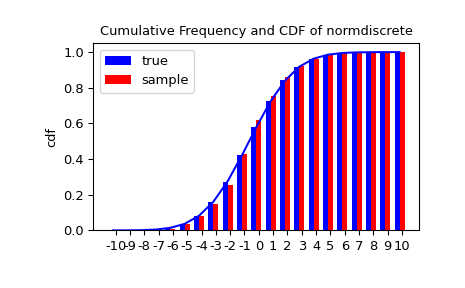
Let’s generate a random sample and compare observed frequencies with the probabilities.
>>> n_sample = 500
>>> rvs = normdiscrete.rvs(size=n_sample)
>>> f, l = np.histogram(rvs, bins=gridlimits)
>>> sfreq = np.vstack([gridint, f, probs*n_sample]).T
>>> print(sfreq)
[[-1.00000000e+01 0.00000000e+00 2.95019349e-02] # random
[-9.00000000e+00 0.00000000e+00 1.32294142e-01]
[-8.00000000e+00 0.00000000e+00 5.06497902e-01]
[-7.00000000e+00 2.00000000e+00 1.65568919e+00]
[-6.00000000e+00 1.00000000e+00 4.62125309e+00]
[-5.00000000e+00 9.00000000e+00 1.10137298e+01]
[-4.00000000e+00 2.60000000e+01 2.24137683e+01]
[-3.00000000e+00 3.70000000e+01 3.89503370e+01]
[-2.00000000e+00 5.10000000e+01 5.78004747e+01]
[-1.00000000e+00 7.10000000e+01 7.32455414e+01]
[ 0.00000000e+00 7.40000000e+01 7.92618251e+01]
[ 1.00000000e+00 8.90000000e+01 7.32455414e+01]
[ 2.00000000e+00 5.50000000e+01 5.78004747e+01]
[ 3.00000000e+00 5.00000000e+01 3.89503370e+01]
[ 4.00000000e+00 1.70000000e+01 2.24137683e+01]
[ 5.00000000e+00 1.10000000e+01 1.10137298e+01]
[ 6.00000000e+00 4.00000000e+00 4.62125309e+00]
[ 7.00000000e+00 3.00000000e+00 1.65568919e+00]
[ 8.00000000e+00 0.00000000e+00 5.06497902e-01]
[ 9.00000000e+00 0.00000000e+00 1.32294142e-01]
[ 1.00000000e+01 0.00000000e+00 2.95019349e-02]]
Next, we can use a chi-squared test, scipy.stats.chisquare, to test the null
hypothesis that the sample is distributed according to our norm-discrete
distribution.
The test requires that there are a minimum number of observations in each bin. We combine the tail bins into larger bins so that they contain enough observations.
>>> f2 = np.hstack([f[:5].sum(), f[5:-5], f[-5:].sum()])
>>> p2 = np.hstack([probs[:5].sum(), probs[5:-5], probs[-5:].sum()])
>>> ch2, pval = stats.chisquare(f2, p2*n_sample)
>>> print('chisquare for normdiscrete: chi2 = %6.3f pvalue = %6.4f' % (ch2, pval))
chisquare for normdiscrete: chi2 = 12.466 pvalue = 0.4090 # random
Conceptually, the test statistic chi2 is sensitive to deviations between
the frequencies of observations and their expected frequencies under the
null hypothesis. The p-value is the probability of drawing samples from the
hypothesized distribution that would produce a statistic value more extreme than
the one we observed. Our statistic value is not very high; in fact, there is
a 40.9% chance that the statistic would be higher than 12.466 if we were to draw a
sample of the same size from the discrete distribution defined by p2. Therefore,
the test provides little evidence against the null hypothesis that the sample was
drawn from our norm-discrete distribution.