scipy.stats.gausshyper¶
-
scipy.stats.gausshyper= <scipy.stats._continuous_distns.gausshyper_gen object>[source]¶ A Gauss hypergeometric continuous random variable.
As an instance of the
rv_continuousclass,gausshyperobject inherits from it a collection of generic methods (see below for the full list), and completes them with details specific for this particular distribution.Notes
The probability density function for
gausshyperis:\[f(x, a, b, c, z) = C x^{a-1} (1-x)^{b-1} (1+zx)^{-c}\]for \(0 \le x \le 1\), \(a > 0\), \(b > 0\), and \(C = \frac{1}{B(a, b) F[2, 1](c, a; a+b; -z)}\). \(F[2, 1]\) is the Gauss hypergeometric function
scipy.special.hyp2f1.gausshypertakes \(a\), \(b\), \(c\) and \(z\) as shape parameters.The probability density above is defined in the “standardized” form. To shift and/or scale the distribution use the
locandscaleparameters. Specifically,gausshyper.pdf(x, a, b, c, z, loc, scale)is identically equivalent togausshyper.pdf(y, a, b, c, z) / scalewithy = (x - loc) / scale.Examples
>>> from scipy.stats import gausshyper >>> import matplotlib.pyplot as plt >>> fig, ax = plt.subplots(1, 1)
Calculate a few first moments:
>>> a, b, c, z = 13.8, 3.12, 2.51, 5.18 >>> mean, var, skew, kurt = gausshyper.stats(a, b, c, z, moments='mvsk')
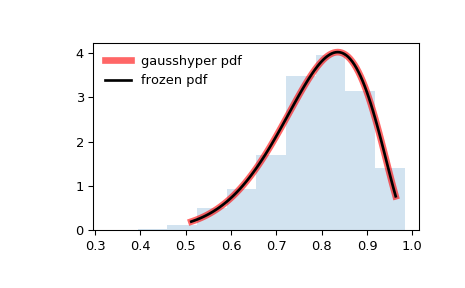
Display the probability density function (
pdf):>>> x = np.linspace(gausshyper.ppf(0.01, a, b, c, z), ... gausshyper.ppf(0.99, a, b, c, z), 100) >>> ax.plot(x, gausshyper.pdf(x, a, b, c, z), ... 'r-', lw=5, alpha=0.6, label='gausshyper pdf')
Alternatively, the distribution object can be called (as a function) to fix the shape, location and scale parameters. This returns a “frozen” RV object holding the given parameters fixed.
Freeze the distribution and display the frozen
pdf:>>> rv = gausshyper(a, b, c, z) >>> ax.plot(x, rv.pdf(x), 'k-', lw=2, label='frozen pdf')
Check accuracy of
cdfandppf:>>> vals = gausshyper.ppf([0.001, 0.5, 0.999], a, b, c, z) >>> np.allclose([0.001, 0.5, 0.999], gausshyper.cdf(vals, a, b, c, z)) True
Generate random numbers:
>>> r = gausshyper.rvs(a, b, c, z, size=1000)
And compare the histogram:
>>> ax.hist(r, density=True, histtype='stepfilled', alpha=0.2) >>> ax.legend(loc='best', frameon=False) >>> plt.show()
Methods
rvs(a, b, c, z, loc=0, scale=1, size=1, random_state=None) Random variates. pdf(x, a, b, c, z, loc=0, scale=1) Probability density function. logpdf(x, a, b, c, z, loc=0, scale=1) Log of the probability density function. cdf(x, a, b, c, z, loc=0, scale=1) Cumulative distribution function. logcdf(x, a, b, c, z, loc=0, scale=1) Log of the cumulative distribution function. sf(x, a, b, c, z, loc=0, scale=1) Survival function (also defined as 1 - cdf, but sf is sometimes more accurate).logsf(x, a, b, c, z, loc=0, scale=1) Log of the survival function. ppf(q, a, b, c, z, loc=0, scale=1) Percent point function (inverse of cdf— percentiles).isf(q, a, b, c, z, loc=0, scale=1) Inverse survival function (inverse of sf).moment(n, a, b, c, z, loc=0, scale=1) Non-central moment of order n stats(a, b, c, z, loc=0, scale=1, moments=’mv’) Mean(‘m’), variance(‘v’), skew(‘s’), and/or kurtosis(‘k’). entropy(a, b, c, z, loc=0, scale=1) (Differential) entropy of the RV. fit(data, a, b, c, z, loc=0, scale=1) Parameter estimates for generic data. expect(func, args=(a, b, c, z), loc=0, scale=1, lb=None, ub=None, conditional=False, **kwds) Expected value of a function (of one argument) with respect to the distribution. median(a, b, c, z, loc=0, scale=1) Median of the distribution. mean(a, b, c, z, loc=0, scale=1) Mean of the distribution. var(a, b, c, z, loc=0, scale=1) Variance of the distribution. std(a, b, c, z, loc=0, scale=1) Standard deviation of the distribution. interval(alpha, a, b, c, z, loc=0, scale=1) Endpoints of the range that contains alpha percent of the distribution
