scipy.stats.nct¶
- scipy.stats.nct = <scipy.stats._continuous_distns.nct_gen object at 0x7f4758331710>[source]¶
A non-central Student’s T continuous random variable.
Continuous random variables are defined from a standard form and may require some shape parameters to complete its specification. Any optional keyword parameters can be passed to the methods of the RV object as given below:
Parameters: x : array_like
quantiles
q : array_like
lower or upper tail probability
df, nc : array_like
shape parameters
loc : array_like, optional
location parameter (default=0)
scale : array_like, optional
scale parameter (default=1)
size : int or tuple of ints, optional
shape of random variates (default computed from input arguments )
moments : str, optional
composed of letters [‘mvsk’] specifying which moments to compute where ‘m’ = mean, ‘v’ = variance, ‘s’ = (Fisher’s) skew and ‘k’ = (Fisher’s) kurtosis. (default=’mv’)
Alternatively, the object may be called (as a function) to fix the shape,
location, and scale parameters returning a “frozen” continuous RV object:
rv = nct(df, nc, loc=0, scale=1)
- Frozen RV object with the same methods but holding the given shape, location, and scale fixed.
Notes
The probability density function for nct is:
df**(df/2) * gamma(df+1) nct.pdf(x, df, nc) = ---------------------------------------------------- 2**df*exp(nc**2/2) * (df+x**2)**(df/2) * gamma(df/2)for df > 0.
Examples
>>> from scipy.stats import nct >>> import matplotlib.pyplot as plt >>> fig, ax = plt.subplots(1, 1)
Calculate a few first moments:
>>> df, nc = 14, 0.240450313312 >>> mean, var, skew, kurt = nct.stats(df, nc, moments='mvsk')
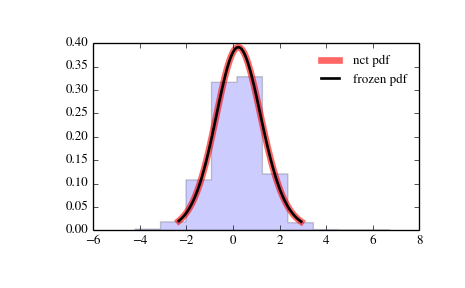
Display the probability density function (pdf):
>>> x = np.linspace(nct.ppf(0.01, df, nc), ... nct.ppf(0.99, df, nc), 100) >>> ax.plot(x, nct.pdf(x, df, nc), ... 'r-', lw=5, alpha=0.6, label='nct pdf')
Alternatively, freeze the distribution and display the frozen pdf:
>>> rv = nct(df, nc) >>> ax.plot(x, rv.pdf(x), 'k-', lw=2, label='frozen pdf')
Check accuracy of cdf and ppf:
>>> vals = nct.ppf([0.001, 0.5, 0.999], df, nc) >>> np.allclose([0.001, 0.5, 0.999], nct.cdf(vals, df, nc)) True
Generate random numbers:
>>> r = nct.rvs(df, nc, size=1000)
And compare the histogram:
>>> ax.hist(r, normed=True, histtype='stepfilled', alpha=0.2) >>> ax.legend(loc='best', frameon=False) >>> plt.show()
Methods
rvs(df, nc, loc=0, scale=1, size=1) Random variates. pdf(x, df, nc, loc=0, scale=1) Probability density function. logpdf(x, df, nc, loc=0, scale=1) Log of the probability density function. cdf(x, df, nc, loc=0, scale=1) Cumulative density function. logcdf(x, df, nc, loc=0, scale=1) Log of the cumulative density function. sf(x, df, nc, loc=0, scale=1) Survival function (1-cdf — sometimes more accurate). logsf(x, df, nc, loc=0, scale=1) Log of the survival function. ppf(q, df, nc, loc=0, scale=1) Percent point function (inverse of cdf — percentiles). isf(q, df, nc, loc=0, scale=1) Inverse survival function (inverse of sf). moment(n, df, nc, loc=0, scale=1) Non-central moment of order n stats(df, nc, loc=0, scale=1, moments=’mv’) Mean(‘m’), variance(‘v’), skew(‘s’), and/or kurtosis(‘k’). entropy(df, nc, loc=0, scale=1) (Differential) entropy of the RV. fit(data, df, nc, loc=0, scale=1) Parameter estimates for generic data. expect(func, df, nc, loc=0, scale=1, lb=None, ub=None, conditional=False, **kwds) Expected value of a function (of one argument) with respect to the distribution. median(df, nc, loc=0, scale=1) Median of the distribution. mean(df, nc, loc=0, scale=1) Mean of the distribution. var(df, nc, loc=0, scale=1) Variance of the distribution. std(df, nc, loc=0, scale=1) Standard deviation of the distribution. interval(alpha, df, nc, loc=0, scale=1) Endpoints of the range that contains alpha percent of the distribution
