correlate2d#
- scipy.signal.correlate2d(in1, in2, mode='full', boundary='fill', fillvalue=0)[source]#
Cross-correlate two 2-dimensional arrays.
Cross correlate in1 and in2 with output size determined by mode, and boundary conditions determined by boundary and fillvalue.
- Parameters:
- in1array_like
First input.
- in2array_like
Second input. Should have the same number of dimensions as in1.
- modestr {‘full’, ‘valid’, ‘same’}, optional
A string indicating the size of the output:
fullThe output is the full discrete linear cross-correlation of the inputs. (Default)
validThe output consists only of those elements that do not rely on the zero-padding. In ‘valid’ mode, either in1 or in2 must be at least as large as the other in every dimension.
sameThe output is the same size as in1, centered with respect to the ‘full’ output.
- boundarystr {‘fill’, ‘wrap’, ‘symm’}, optional
A flag indicating how to handle boundaries:
fillpad input arrays with fillvalue. (default)
wrapcircular boundary conditions.
symmsymmetrical boundary conditions.
- fillvaluescalar, optional
Value to fill pad input arrays with. Default is 0.
- Returns:
- correlate2dndarray
A 2-dimensional array containing a subset of the discrete linear cross-correlation of in1 with in2.
Notes
When using “same” mode with even-length inputs, the outputs of
correlateandcorrelate2ddiffer: There is a 1-index offset between them.Array API Standard Support
correlate2dhas experimental support for Python Array API Standard compatible backends in addition to NumPy. Please consider testing these features by setting an environment variableSCIPY_ARRAY_API=1and providing CuPy, PyTorch, JAX, or Dask arrays as array arguments. The following combinations of backend and device (or other capability) are supported.Library
CPU
GPU
NumPy
✅
n/a
CuPy
n/a
✅
PyTorch
✅
⛔
JAX
✅
✅
Dask
⚠️ computes graph
n/a
The JAX backend only supports
boundary="fill"andfillvalue=0.See Support for the array API standard for more information.
Examples
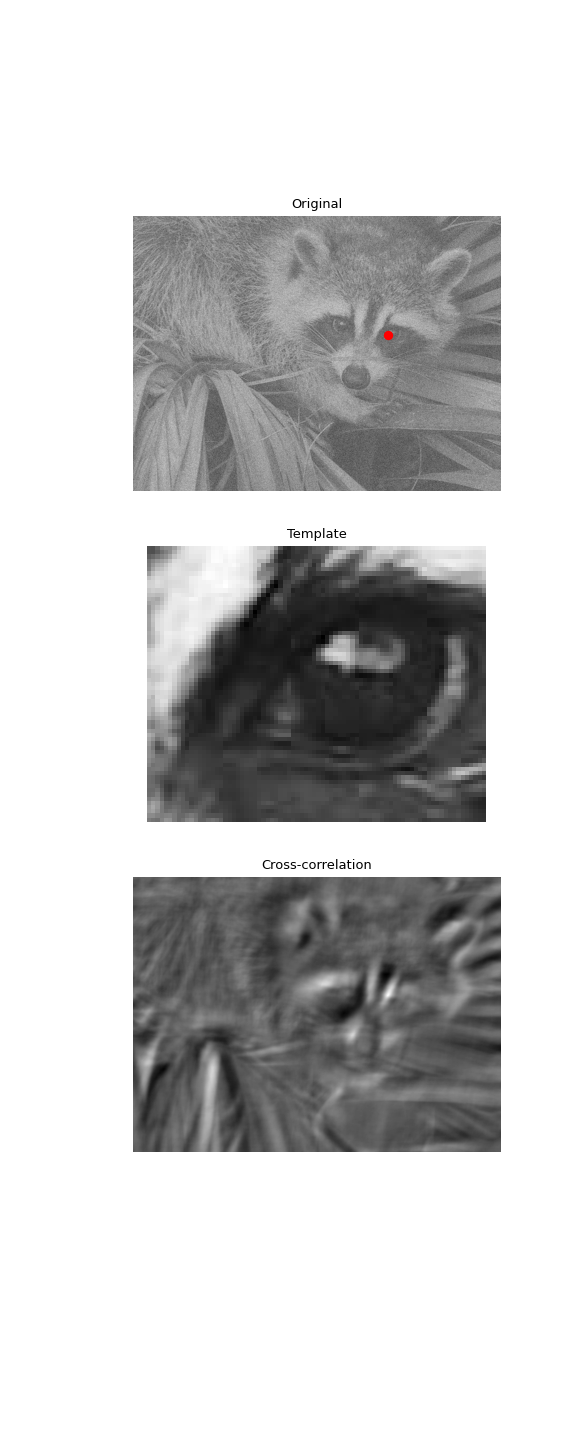
Use 2D cross-correlation to find the location of a template in a noisy image:
>>> import numpy as np >>> from scipy import signal, datasets, ndimage >>> rng = np.random.default_rng() >>> face = datasets.face(gray=True) - datasets.face(gray=True).mean() >>> face = ndimage.zoom(face[30:500, 400:950], 0.5) # extract the face >>> template = np.copy(face[135:165, 140:175]) # right eye >>> template -= template.mean() >>> face = face + rng.standard_normal(face.shape) * 50 # add noise >>> corr = signal.correlate2d(face, template, boundary='symm', mode='same') >>> y, x = np.unravel_index(np.argmax(corr), corr.shape) # find the match
>>> import matplotlib.pyplot as plt >>> fig, (ax_orig, ax_template, ax_corr) = plt.subplots(3, 1, ... figsize=(6, 15)) >>> ax_orig.imshow(face, cmap='gray') >>> ax_orig.set_title('Original') >>> ax_orig.set_axis_off() >>> ax_template.imshow(template, cmap='gray') >>> ax_template.set_title('Template') >>> ax_template.set_axis_off() >>> ax_corr.imshow(corr, cmap='gray') >>> ax_corr.set_title('Cross-correlation') >>> ax_corr.set_axis_off() >>> ax_orig.plot(x, y, 'ro') >>> fig.show()